Reduce sequencing cost with high quality libraries
The combination of low PCR background, low GC bias, and high mapping rate, on-target rate, and coverage uniformity means that very few sequencing reads are wasted on sequencing off-target sequences and non-specific PCR products and primer-dimers. It also means that fewer sequencing reads are required to ensure all targets are covered at a minimally required depth to make confident base calls. Overall, CleanPlex® NGS Panels allow efficient use of sequencing reads so that sequencing can be performed cost-effectively by allowing more samples to be sequenced at a time.
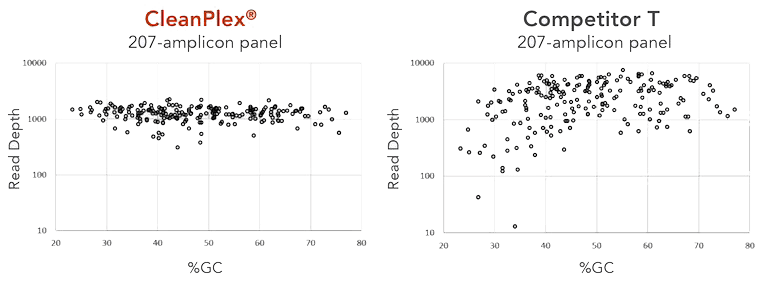
High performance translates to cost-effective sequencing. A 207-amplicon panel was used to generate target-enriched NGS libraries using either the CleanPlex or Competitor T’s library preparation chemistry. Libraries were sequenced and analyzed for coverage uniformity across GC content. The results indicate that for a 207-amplicon panel, 60% less sequencing would be required using CleanPlex, which means 2.5X more samples can be sequenced on a flow cell. To achieve similar data quality, CleanPlex’s mean read depth could be reduced to 600X coverage while Competitor T’s would need to be increased to >1,500X coverage.