Product Description
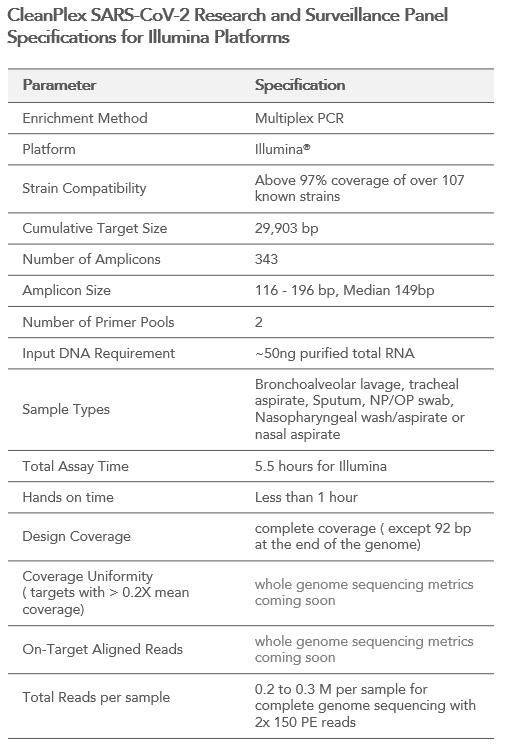
CleanPlex® SARS-CoV-2 Panel for COVID-19 coronavirus detection, tracking, and research via amplicon-based target enrichment for Next Generation Sequencing (NGS) on Illumina sequencing platforms
Paragon Genomics designed a highly multiplexed target enrichment panel covering the entire genome of the SARS-CoV-2 virus (except for 92 bases at the ends). The panel enables complete genome sequencing and epidemiological studies of the new SARS-CoV-2 virus responsible for the COVID-19 pandemic. With our CleanPlex technology, the entire genome of the virus can be amplified from RNA to sequence-ready libraries in 5.5 hours. Two versions of this panel are available for two major sequencing platforms: Illumina and MGI. CleanPlex Technology allows the ease and flexibility to sequence samples for confident infectious disease surveillance and research.
The CleanPlex SARS-CoV-2 FLEX Kit is also available for front-end leading coronavirus research customers and those who may benefit from internal library preparation control.
The CleanPlex SARS-CoV-2 Kit has repeatedly demonstrated consistently high coverage of multiple variants. Read more about how the panel has been used around the globe for surveillance and research.
To ensure that our customers get the most out of their sequencing runs, we’re proud to offer up to 2688 sample multiplexing capability with our combinatorial dual-indexed primers for Illumina sequencing. Additionally, we provide 384 unique-dual indexes, allowing for low variant calling and other in-depth sequencing applications. These exciting additions allow for cost-effective and high-throughput sequencing with our best-in-class CleanPlex targeted sequencing technologies.
Highlights
- Significant Cost Savings
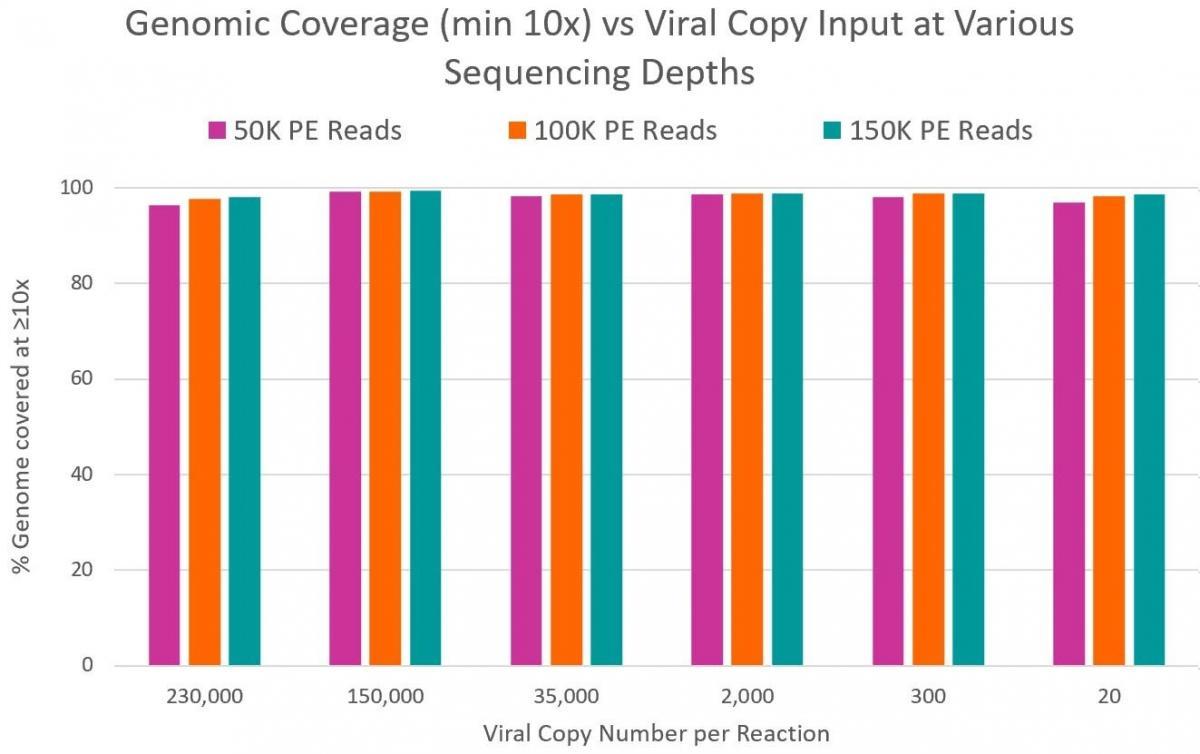
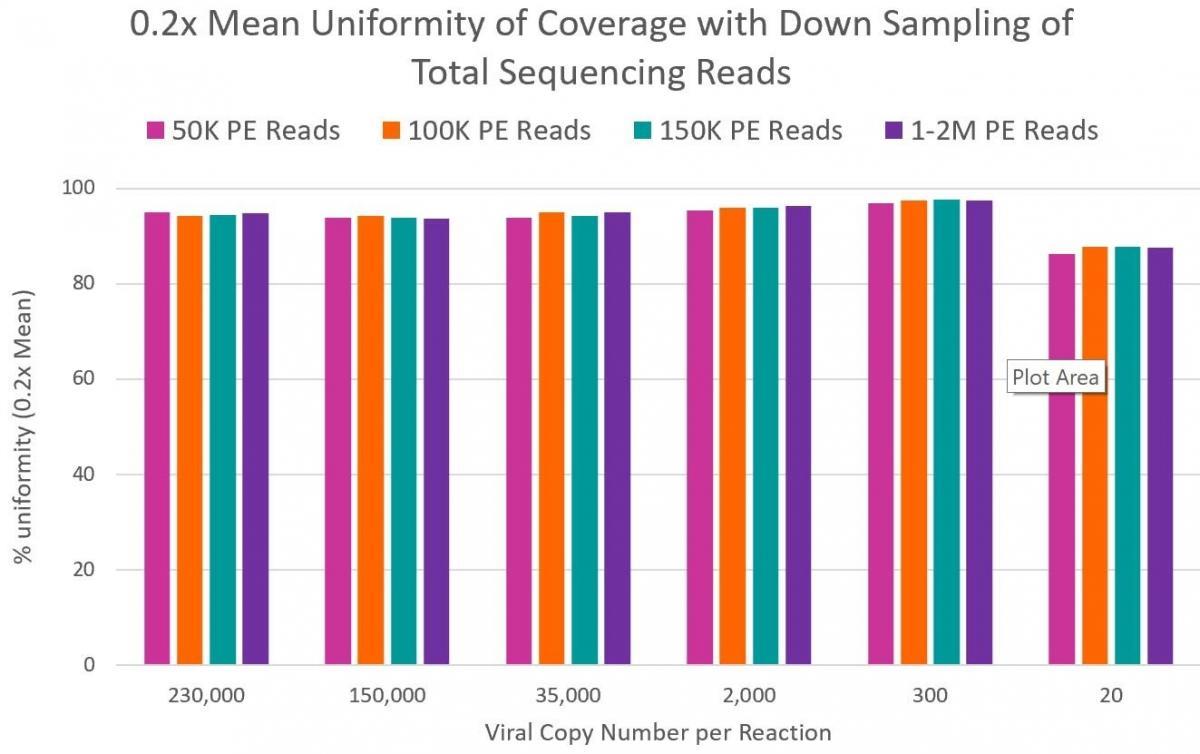
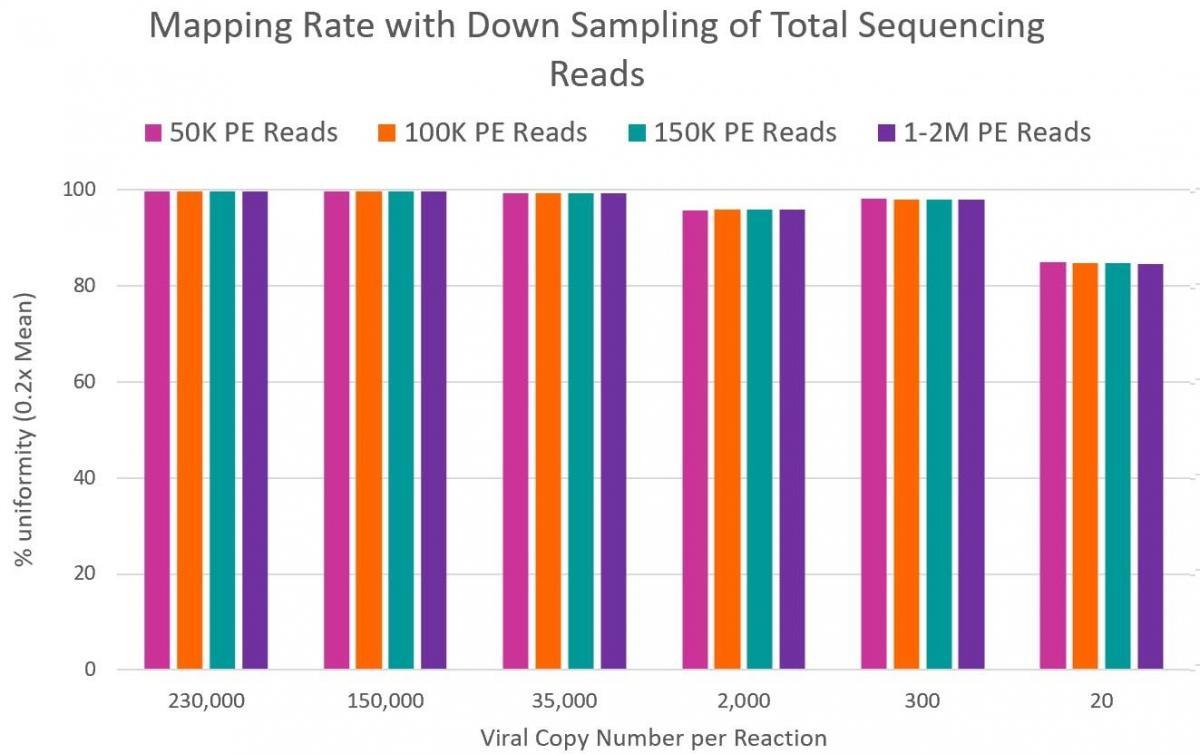
99% reduction in sequencing cost. Only 50K reads per sample required versus 10M reads required by many shotgun metagenomics sequencing methods. - High Coverage of Target Regions
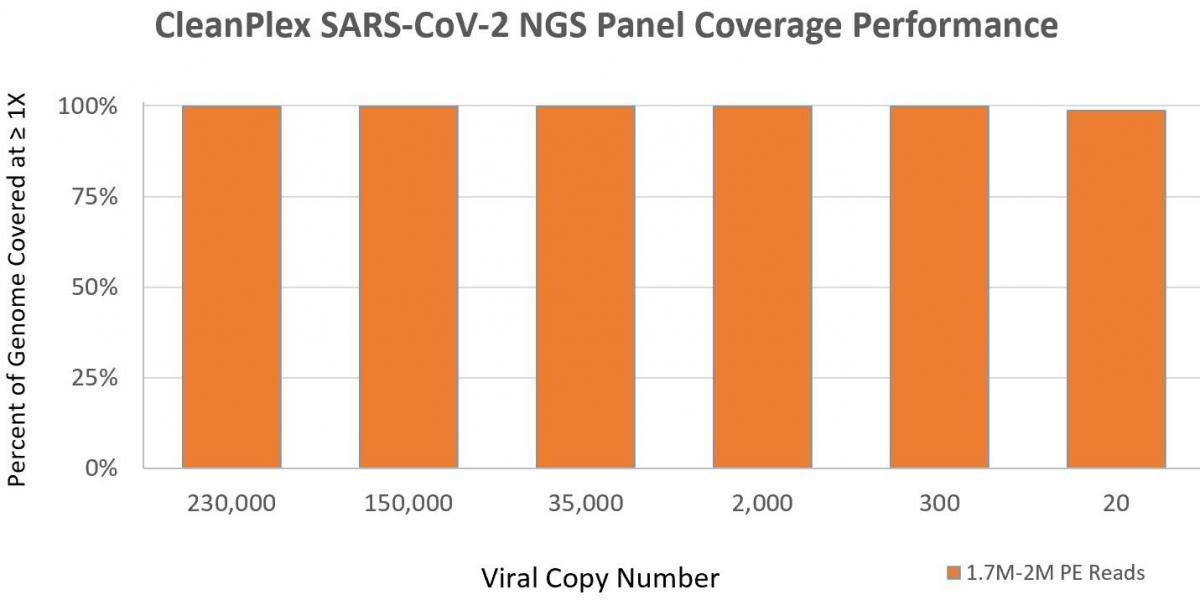
99% coverage of the entire SARS-CoV-2 genome - Sensitive Detection
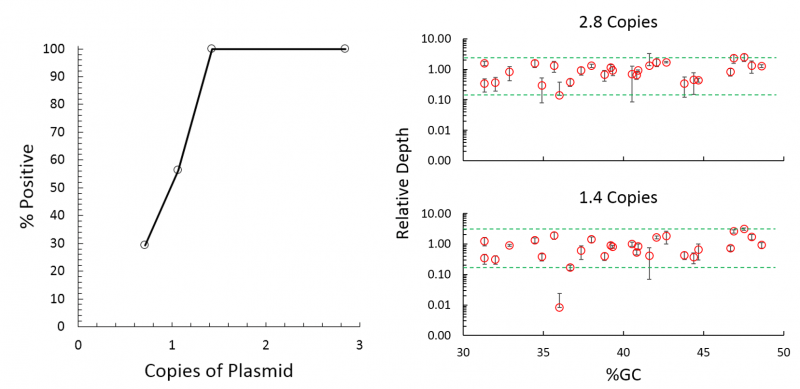
Can detect down to 1 copy with high confidence compared 3-5 copies by RT-qPCR (1) - Fast, Streamlined Workflow
Generate sequencing-ready libraries in just 5.5 hours using a rapid, four-step protocol including RT step with minimal hands-on time. - Superb Performance
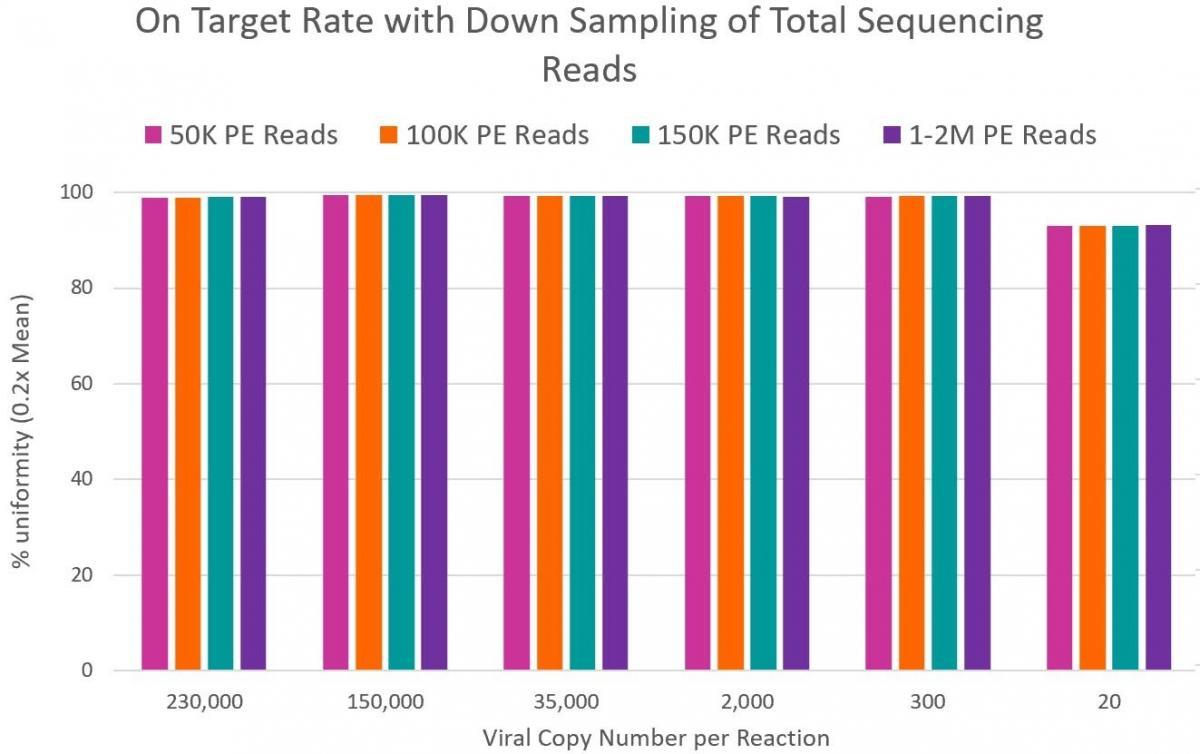
Prepare high-quality NGS libraries with excellent on-target performance using CleanPlex Technology to enable efficient use of sequencing reads. The CleanPlex panels’ on-target rates are usually much higher than those of hybrid captured-based small target enrichment panels.
The CleanPlex SARS-CoV-2 Kit contains the CleanPlex Multiplex PCR Primers and CleanPlex Targeted Library Kit with RT components. CleanPlex Indexed PCR Primers for sample pooling and CleanMag® Magnetic Beads for bead purification and size selection are ordered separately to complete the workflow from input RNA to sequencing-ready NGS libraries.
For a complete solution, please use the following indexed PCR primers and magnetic beads SKUs.
Storage Temperature
Store at -20 °C.
(1) Corman VM et al. Detection of 2019 novel coronavirus (2019-nCoV) by real-time RT-PCR https://www.ncbi.nlm.nih.gov/pmc/articles/PMC6988269/
For Research Use Only. Not for use in diagnostic procedures.