What is Next-Generation Sequencing (NGS)?
Next-Generation Sequencing (NGS) is a massively parallel method to sequence thousands to millions of DNA molecules simultaneously. Since NGS was first introduced in 2005, it has risen to become the predominant DNA sequencing technology and heralded new levels of accessibility for research, industry, and even everyday consumers alike. Most importantly, it has begun to make precision medicine a reality, with clinicians able to sequence a patient’s whole genome, exome or a targeted gene panel cheaply, and prescribe therapies tailored to individual genetic profiles.
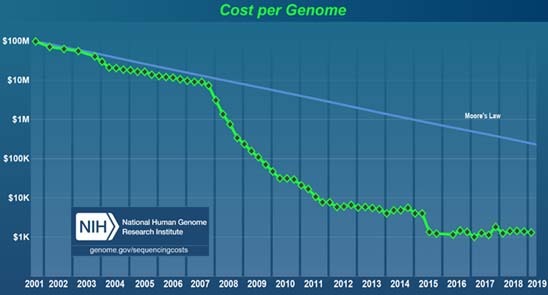
Improvements in DNA sequencing technologies since 1990 have resulted in dramatic reductions in the time and cost of sequencing a single human genome. Today, NGS technology can sequence a human whole genome (more than 3 billion basepairs) within two days at a dramatically lower cost. Below is a chart that illustrates the DNA sequencing cost trend from 2001 – 2019.
What is Targeted Sequencing (Resequencing) / Target Enrichment?
Targeted Sequencing or Resequencing is a method for only sequencing part of a whole genome or regions of interest without sequencing the entire genome of a sample. In order to only focus on specific or clinically relevant regions of a genome or DNA sample, it requires a pre-sequencing DNA preparation step called Target Enrichment where target DNA sequences are either directly amplified (amplicon or multiplex PCR-based) or captured (hybrid capture-based) and then subsequently sequenced using DNA sequencers.
Why Target Enrichment?
Whole genome sequencing (WGS) and its corresponding whole genome amplification (WGA) method are more suitable for research and discovery types of applications, while targeted sequencing and target enrichment are essential to many fast-growing clinical and industrial applications where cost and speed are more important.
One of the challenges the genomics community faces is the continued acquisition of large amounts of sequencing raw data that is yet to be fully and successfully translated and interpreted to help advance research, diagnose and cure diseases on a wider scale1. Even with current reduced sequencing costs, a whole genome sequencing approach can be practically used only in specific scenarios such as basic research, population genetics or rare disease detection. A focused or targeted approach would be more appropriate for understanding disease progression and guiding therapy selection in clinical setting or massively screening DNA samples in industrial applications. In addition, sequencing an entire genome or exome can be prohibitively expensive in terms of laboratory operations and bioinformatics infrastructure for storing and processing large amounts of data2,3. Therefore, target enrichment has become vital for the continued progress of precision medicine and research.
The following table compares WGS and targeted sequencing in terms of sequencing and library preparation reagent costs. In addition to reagent costs, bioinformatics analysis time and cost shall also be taken into account when considering the suitable methods for your applications.
| Comparison of WGS and Targeted Sequencing Methods | |||
| WGS | Targeted Sequencing | ||
| Human Whole Genome | Human Whole Exome (50Mb) | Targeted Panel (1Mb) | |
| Target Region Size (basepair) | 3 x 109 | 5 x 107 | 1 x 106 |
| Depth of Coverage | 30X | 100X | 1000X |
| Number of Samples per Sequencing Run* | 8 | 145 | 725 |
| Cost of Sequencing Reagent per Sample** | $1,500 | $100 | $15 |
| Cost of Target Enrichment per Sample*** | N/A | $250 | $100 |
| Total Cost per Sample (excluding bioinformatics) | $1,500 | $350 | $115 |
* Based on Illumina HiSeq 2500 System dual flow cell high output mode
** Based on Illumina HiSeq SBS V4 cost
*** Based on average target enrichment and library preparation kit prices
The clear benefits of targeted resequencing are driving the adoption of NGS in areas such as translational research, clinical diagnostics and industrial applications while the value also shifts from sequencing reagents to target enrichment reagents on a per sample basis. For a small targeted sequencing panel covering for example only 100 kb region of interest, most of the cost is incurred at the library preparation and target enrichment steps while the cost of sequencing reagents is negligible for each given sample.
What are major applications of Targeted Sequencing and Target Enrichment?
Currently, targeted sequencing and target enrichment have been applied to many areas from basic research to clinical diagnostics and applied markets. Below is a list of major applications.
- Cancer
- Cancer research and diagnostics
- Tumor profiling
- Cancer liquid biopsy
- Immuno-Oncology
- Minimal residual disease testing
- Circulating Tumor Cell (CTC) analysis
- Reproductive Health
- Carrier screening
- Non-invasive prenatal testing
- Preimplantation genetic diagnosis (PGD)
- Newborn screening
- Cardiovascular Disease Testing
- Infectious Disease Testing and Surveillance
- Transplant Genomics / HLA typing
- Inherited Disease Testing
- Metabolic and Immune Disorders
- Neurological Disease Testing
- Companion Diagnostics
- Applied / Industrial Applications
- Agrigenomics / molecular breeding
- Food safety and animal health
- Forensics
- Environmental research
The common techniques employed for target enrichment are mainly based on multiplex PCR, hybridization (hybrid capture probe), molecular inversion probe (originated from padlock probe), or low- / single-plex PCR in microfluidic compartments. Here we will mainly discuss the two most commonly used NGS target enrichment strategies: amplicon- or multiplex PCR-based enrichment and hybrid capture-based enrichment2,3.
In general, multiplex PCR-based method is faster, easier and cheaper than hybridization-based alternatives and so more suited for quick gene panel testing and production-scale applications such as clinical diagnostics and industrial genomic screening. Multiplex PCR-based enrichment also allows for dealing with low DNA input which is difficult for hybrid captured based technology. On the other hand, hybrid capture-based method can interrogate very large target region up to human whole exome and is generally better at detecting structural variations such as novel gene fusions, etc., so it is more suitable for research and discovery projects. However, capture-based method suffers low on-target rate on smaller panels due to its inherent lower specificity of hybridization probes. Due to the above reasons, scientists and clinicians tend to employ multiplex PCR for small (less than 1Mb target region) or hotspot panels for the detection of single nucleotide polymorphisms (SNPs) and/or small insertions/deletions and hybrid capture for large (0.5Mb – 50Mb target region) panels for the detection of SNPs, fusion genes, copy number variations (CNV), etc.
Below is a comparison of typical workflows of Amplicon-based target enrichment and hybrid capture-based target enrichment.
| Amplicon- or PCR-based target enrichment workflow | Hybrid Capture-based target enrichment workflow |
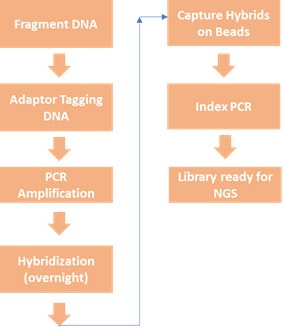 |
Below is a summary of the pros and cons of both methods.
Note: Some comments on Amplicon-based target enrichment do not apply to Paragon Genomics’ CleanPlex technology as it overcomes many of the drawbacks of traditional amplicon-based methods. This will be discussed in the next section.
| Traditional Amplicon- or PCR-based Target Enrichment | Hybrid Capture-based Target Enrichment |
| Pros | |
| Requires lower input of DNA (1-10 ng) | Can support large panel size (up to human whole exome) |
| Shorter and easy workflow (3-6 hours) | Can detect some novel structural variations such as novel gene fusions |
| High on-target rate (>95%) | has better assay uniformity in general |
| No special equipment required (e.g. no need of DNA fragmentation) | Custom assay probes are easier to design |
| Better performance on difficult clinical samples such as FFPE tissue DNA | |
| Lower reagent and consumable (pipette tips) costs | |
| Cons | |
| PCR amplification bias resulting in lower assay uniformity, especially evident for large panels | Needs to fragment DNA first |
| Non-specific PCR background noise (e.g. primer dimer) can be high, especially evident for large panels involving over 1,000 amplicons in a single pool | Requires higher input DNA (>10 ng) |
| Difficult to design a large number of multiplex PCR primer pairs that are compatible in a single pool with minimal interaction | Longer and laborious workflow (>10 hours) |
| Low on-target rate for small panels and mixed samples (e.g. detection of bacteria or viruses in human DNA background); some as low as 10-20% | |
| Higher reagent and consumable (pipette tips) costs | |
| Application Limitation | |
| Novel fusion detection (cannot design primers for unknown fusion breakpoints) | TCR/BCR sequencing (Application doesn’t allow for fragmenting genomic DNA due to the need to preserve TCR/BCR variable regions’ integrity) |
| Whole Exome Sequencing (target region can be too large and uniformity will suffer) | Pathogen detection on mixed samples with varying DNA inputs (e.g. TB patient samples) |
CleanPlex® Technology – a novel ultra-high multiplex PCR-based target enrichment method that bridges the gap between traditional multiplex PCR and hybrid capture
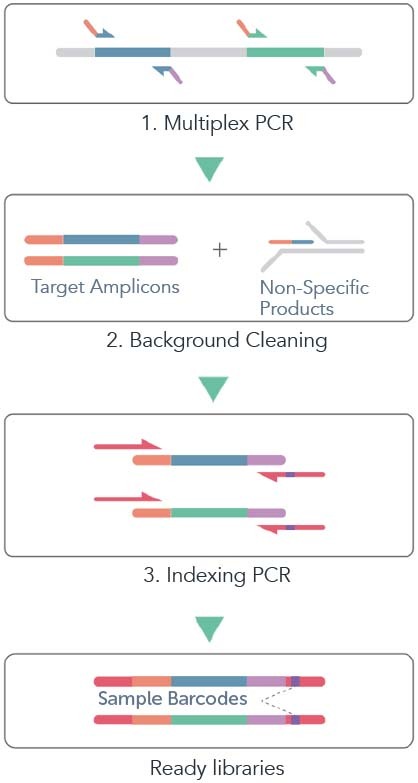
CleanPlex® technology is a novel amplicon-based NGS target enrichment technology developed by Paragon Genomics. The technology inherits major advantages associated with traditional multiplex PCR methods while overcoming their key shortcomings such as PCR background noise, scalability (panel size), uniformity and limitation on detecting novel fusion genes. The schematic below shows an example workflow of CleanPlex Technology.
Overcoming Background Noise
First, the targeted regions of a genome or DNA sample are amplified by well-designed multiplex PCR primers with overhanging tails being partial adaptor sequences compatible with corresponding DNA sequencers, resulting in both target amplicons and non-specific PCR products including primer dimers.
Traditionally, when the panel size is large (e.g. more than 1,000 amplicons in a single pool), the non-specific PCR products can be overwhelming and significantly affect the downstream steps if there is no measure to remove them. Some amplicon-based methods utilize bead purification and size selection to remove smaller DNA fragments such as primer dimers. However, some complicated non-specific PCR products with sizes similar to the lengths of target amplicons and its resulting libraries can be difficult to remove using just size selection. The following Bioanalyzer trace shows significant background noise around a target library of 300bp.
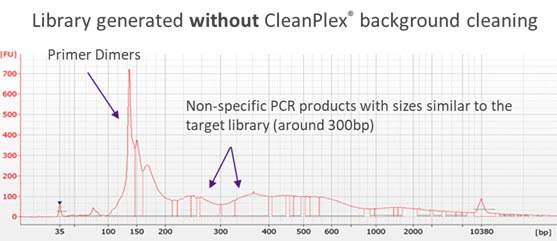
CleanPlex overcomes this drawback with an innovative and patented enzymatic background cleaning step that removes non-specific PCR products including both primer dimers and more complicated and longer nonspecific PCR artifacts, resulting in very pure target libraries. The following Bioanalyzer trace shows the effect of CleanPlex background cleaning technology.
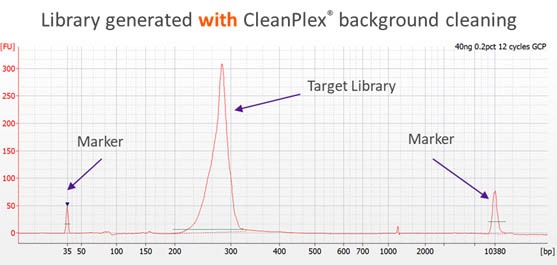
Subsequently, sample barcodes (for sample pooling purpose) are added by an indexing PCR step to get sequencing-ready libraries. The whole workflow only takes 3 hours and minimal hands-on time.
Overcoming Scalability or Panel Size Limitation
Due to its background cleaning technology, CleanPlex can easily break the panel size limit of traditional multiplex PCR and amplify more than 20,000 targets in a single panel. The following amplification plot shows the GC% of each target amplicon vs its sequencing depth for a 27,000-amplicon panel.
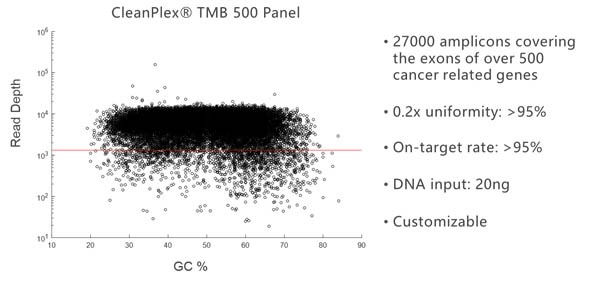
Overcoming GC Bias and Uniformity Issues
The figure below compares CleanPlex and another well-known amplicon-based method in terms of GC bias and amplification uniformity. Two panels target the same regions of a few cancer-related genes. The competitor method obviously has GC bias around low GC region and an overall low amplification uniformity across the spectrum of the panel. On the contrary, CleanPlex can amplify all target regions evenly. The result of this is that users of the competitor’s panel have to raise the average sequencing depth by 100% to achieve similar variant calling quality, therefore doubling the sequencing depth and cost required for CleanPlex panel.
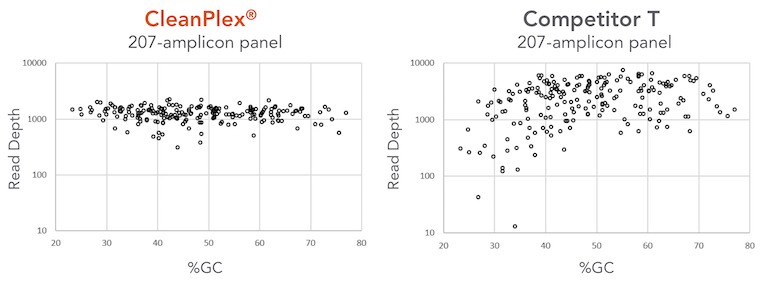
Pushing the limit of LOD
Our partner RareCyte, Inc has successfully demonstrated that CleanPlex technology can directly amplify DNA from a single cell (only ~6 pg of DNA), specifically circulating tumor cells isolated from cancer patient blood.
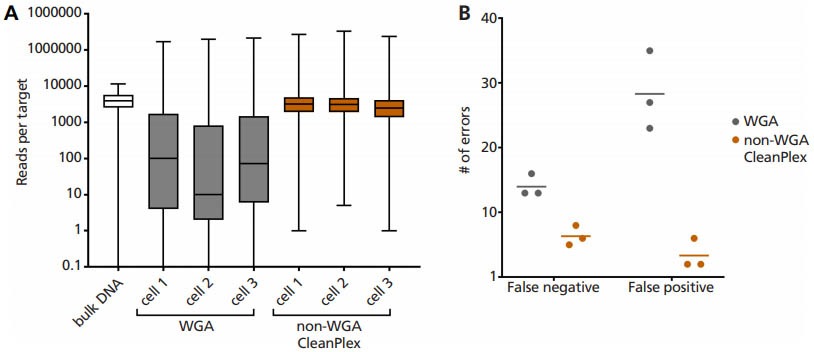
Single cell lysate is input as template into Paragon Genomics’ CleanPlex OncoZoom Panel, with modified primer concentration, PCR cycle number, and clean-up steps to compensate for low input DNA concentration. Using this non-WGA method vastly improves: (A) coverage uniformity, and incidence of (B) false negative and false positive errors, when compared to single cell WGA products.
Overcoming limitation on Detecting Novel Fusion Genes
CleanPlex OmniFusion Technology (to be launched soon) leverages template switching and single primer amplification methods to detect novel fusion genes.
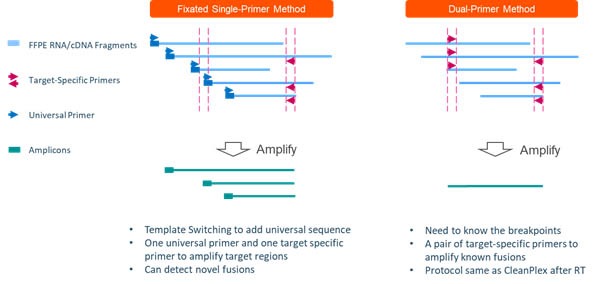
In summary, CleanPlex Target Enrichment Technology leverages the speed, sensitivity, workflow and cost advantages of multiplex PCR for NGS target enrichment while overcoming its key drawbacks, therefore bridging the gap between traditional amplicon-based target enrichment method and hybrid captured-based approach.
| Traditional Amplicon- or PCR-based Target Enrichment | CleanPlex Improvements |
| Pros | |
| Requires lower input of DNA (1-10 ng) | CleanPlex can directly amplify single cell DNA (6 pg) |
| Shorter and easy workflow (3-6 hours) | Advantages maintained |
| High on-target rate (>95%) | Advantages maintained |
| No special equipment required (e.g. no need of DNA fragmentation) | Advantages maintained |
| Better performance on difficult clinical samples such as FFPE tissue DNA | Advantages maintained |
| Lower reagent and consumable (pipette tips) costs | Advantages maintained |
| Cons | |
| PCR amplification bias resulting in lower assay uniformity, especially evident for large panels | Can achieve more than 95% uniformity (0.2x) even for large panels involving over 20,000 amplicons |
| Non-specific PCR background noise (e.g. primer dimer) can be high, especially evident for large panels involving over 1,000 amplicons in a single pool | Proprietary CleanPlex background cleaning technology can effectively remove background noise regardless of panel sizes |
| Difficult to design a large number of multiplex PCR primer pairs that are compatible in a single pool with minimal interaction | ParagonDesigner™ algorithm takes into account multiple primer design factors and ensures best performance for even very large panels |
| Application Limitation | |
| Novel fusion detection (cannot design primers for unknown fusion breakpoints) | CleanPlex OmniFusion single primer technology enables detection of novel fusions |
| Whole Exome Sequencing (target region can be too large and uniformity will suffer) | CleanPlex is the most scalable and uniform multiplex-PCR based technology and will be able to amplify whole exome. |
References:
- Human genome sequencing in health and disease. Gonzag-Jauregui C. et al. (2012) Annu. Rev. Med. 63:35–61
- Target-enrichment strategies for next-generation sequencing. Mamanova L. et al. (2010) Nat. Methods. 7(2):111–118
- Overview of target enrichment strategies. Kozarewa I. et al. (2015) Curr. Protoc. Mol. Biol.112: 7.21.1–7.21.23
- Advances in clinical next-generation sequencing: target enrichment and sequencing technologies. Ballester L.Y. et al. (2016) Expert Rev. Mol. Diagn. 16(3): 357–372
- Evaluation of hybridization capture versus amplicon-based methods for whole-exome sequencing. Samorodnitsky E et al. (2015) Hum. Mutat. 36(9):903-14
Enter your email below to schedule a free consultation with our PhD-level expert scientists to learn how our best-in-class custom panel design services can help you advance your work.
At Paragon Genomics, we take your privacy and data very seriously. You can review our Privacy Policy online or download the PDF.