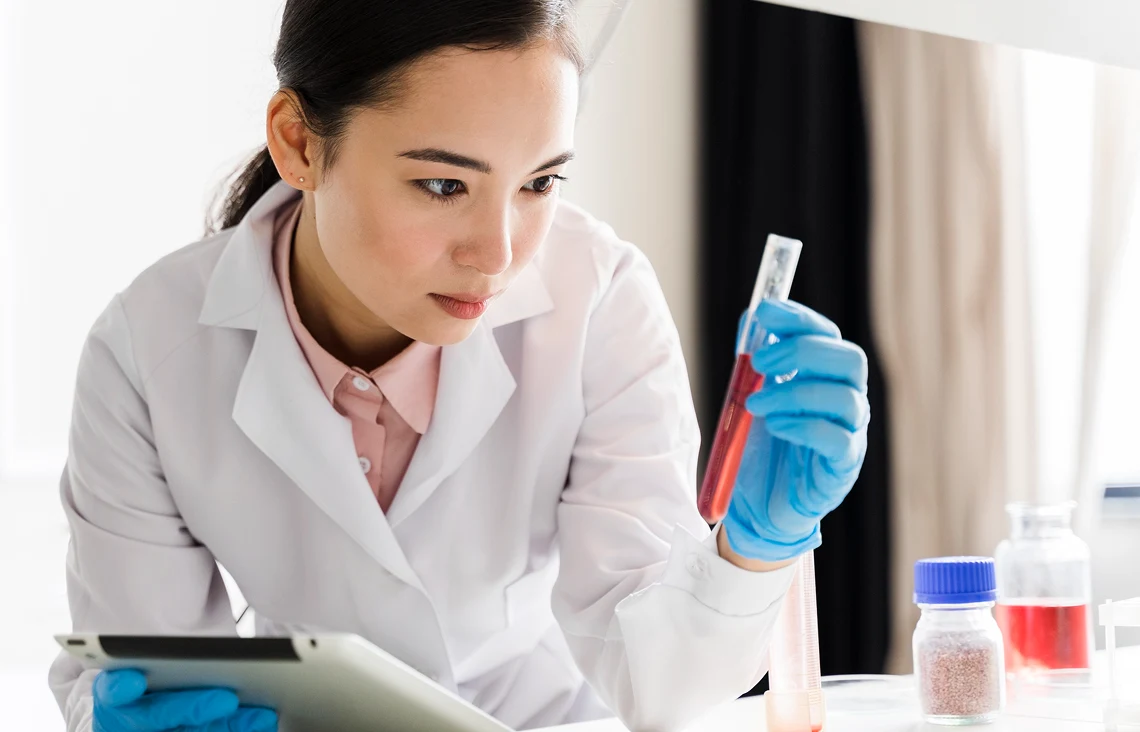
Targeted Sequencing
CleanPlex® technology drives targeted sequencing by focusing only on specific regions of the genome, offering a faster and more cost-effective alternative to whole genome sequencing. By enriching and sequencing areas of interest, CleanPlex® generates actionable data with less complexity, helping accelerate advances in clinical diagnostics, infectious disease monitoring, and precision oncology.
Advantages of Targeted Sequencing with CleanPlex®:
- Faster turnaround times compared to whole genome sequencing
- Lower costs for sequencing and data analysis
- Focused data generation for easier interpretation and clinical relevance
- Ideal for large-scale screening and therapy selection
Amplicon Sequencing
CleanPlex® technology powers amplicon sequencing by using highly efficient multiplex PCR to selectively amplify target regions. This method delivers high sensitivity, works with low DNA input, and produces rapid results, making it a preferred solution for applications such as viral detection, industrial genomic screening, and mutation analysis.
Advantages of Amplicon Sequencing with CleanPlex®:
- Faster, easier, and more affordable than hybrid capture methods
- High sensitivity even with low DNA or RNA input
- Quick and scalable workflow for production-scale applications
- Strong performance for detecting rare mutations, SNPs, insertions, deletions, and gene fusions
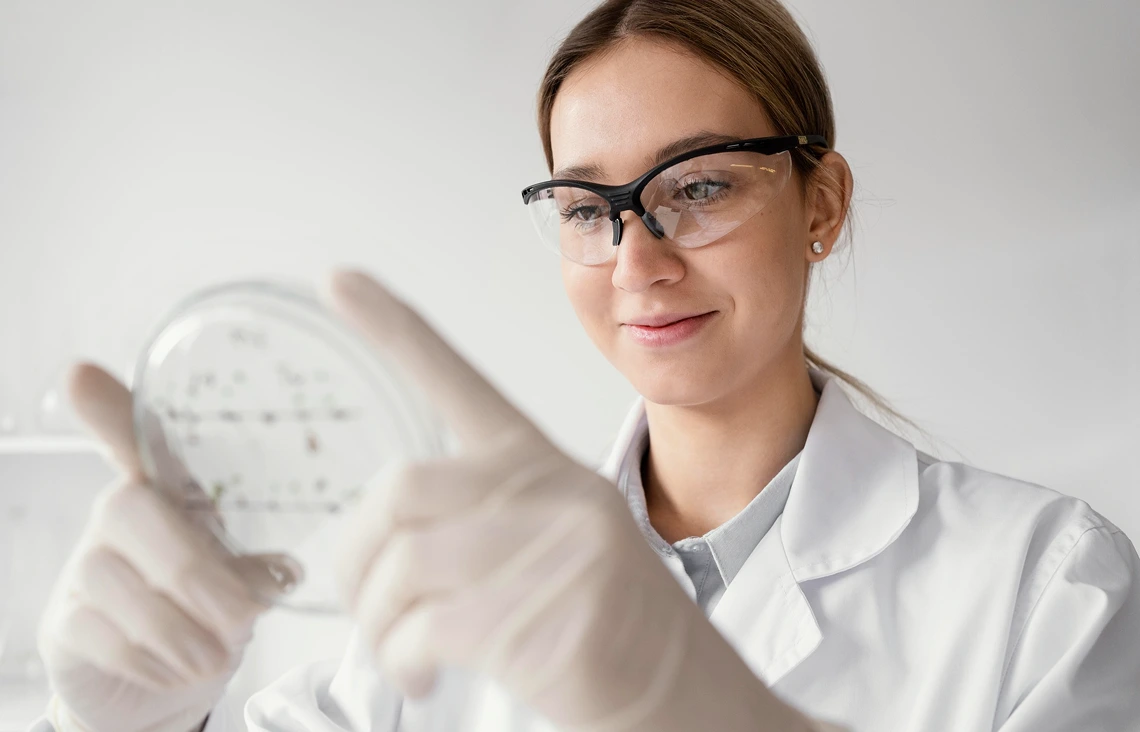

Target Enrichment
Target enrichment prepares DNA samples by amplifying or capturing regions of interest before sequencing. It forms the foundation of targeted sequencing, helping researchers avoid the cost and complexity of sequencing an entire genome. Paragon Genomics’ CleanPlex® Target Enrichment stands out for its rapid three-step workflow, high on-target rates, and minimal hands-on time.
Advantages of Target Enrichment with CleanPlex®:
- Sequencing-ready libraries prepared in under 3 hours
- Reduced overall sequencing costs through high multiplexing capability
- Highly sensitive detection with low limits of detection
- Outstanding DNA enrichment performance with high mapping rates and uniformity
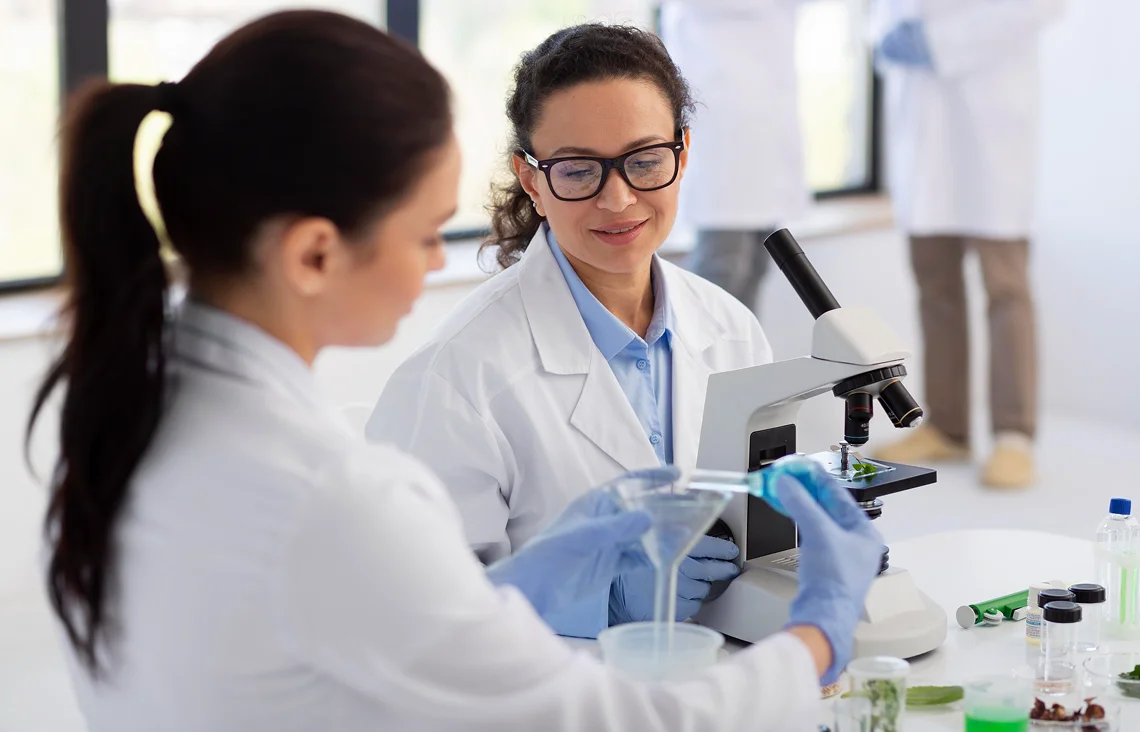

Sequencing Platform Compatibility
Paragon Genomics designs its CleanPlex® solutions to work seamlessly with any and all sequencing platforms like Illumina and Ion Torrent. This compatibility gives labs flexibility to match their library preparation kits to existing workflows, budget needs, and project timelines, while maintaining high performance across different applications.
Advantages of Using CleanPlex® for Sequencing Platform Compatibility:
- Sequencer Agnostic: Can manufacture for any sequencer
- Broad platform support for a variety of research and clinical applications
- Easy integration into existing lab setups
- Faster time-to-results with streamlined compatibility
- Strong performance across diverse fields like oncology, infectious disease, agrigenomics, and precision medicine